Analysis of β-casein gene for A1 and A2 allelic variants in Holstein-Local crossbred cattle of Bangladesh using allele-specific PCR
Abstract
β-casein (CSN2) is one of the important bovine milk proteins synthesized by β-casein gene. The aim of this study was to investigate the A1 and A2 allelic variants of CSN2 gene in Holstein-Friesian and Local (HF×L) crossbred cattle of Bangladesh. Allele specific polymerase chain reaction (AS-PCR) was carried out for genotyping of allelic variants. A total of 390 blood samples of HF×L crossbred cattle was included from government, military, and private dairy farms. Six different crossbred genotypes were identified in the studied population such as 50%HF×50%L, 62.5%HF×37.5%L, 68.75%HF×31.25%L, 75%HF×25%L, 81.25%HF×18.75%L, and 87.5%HF×12.5%L. The mean frequencies of the A1A1, A1A2, and A2A2 genotypes were 0.11, 0.48, and 0.41, respectively, while the corresponding frequencies of the A1 and A2 alleles were 0.34 and 0.66, respectively. The frequency of desirable A2A2 genotype reduced from 0.60 (50%HF×50%L) to 0.38 (87.5%HF×12.5%L) with the advancement of exotic HF inheritance from 50 to 87.5%. In contrast, the frequency of A1A1 and A1A2 genotypes increased proportionately with the progression of HF inheritance level. The undesirable A1A1 genotype frequency was found highest in the HF crossbreds of private dairy farms (0.13) compared to government (0.08) and military dairy farm (0.07) that signifies the availability of upper graded cows in the private dairy farms. Altogether, this study provides current scenario of A1 and A2 allelic variants of HF×L crossbreds of Bangladesh that could be utilized to design suitable breeding plans in crossbreeding programs for production of premium quality milk.
INTRODUCTION
Proteins are an essential component of milk, including 80% of milk casein and the remaining 20% of whey proteins (α-lactalbumin and β-lactoglobulin). Milk casein is made up of four different types of casein: α-casein S1 (CSN1S1), α-casein S2 (CSN1S2), β-casein (CSN2) and κ-casein (CSN3) [1]. The second most abundant protein in cow's milk is CSN2 which makes up 25–30% of the total protein content and consists of 209 amino acids [2]. The bovine β-casein gene belongs to a cluster of four casein family which are located on chromosome BTA6 [1,3]. Importantly, CSN2 gene functions as a molecular marker for milk production traits in dairy cattle and plays important roles in the quantity or quality of cheese and the renneting properties of milk [4, 5]. Among the various kinds of milk casein, the most polymorphic milk protein gene in cattle is CSN2. The fifteen genetic variants of the CSN2 gene, which are caused by numerous amino acid substitutions in the CSN2 protein that alter the protein's properties, are A1, A2, A3, B, C, D, E, F, G, H1, H2, I, J, K, and L [6,7]. Among the CSN2 variants, the A1 and A2 types are most common variants while B is less common, and A3 and C are rare [1,5,8]. It is considered that A2 CSN2 is the original casein protein because it was present before the point mutation caused the advent of A1 CSN2 in European cattle (Bos taurus) populations several thousand years ago [3,9].
One nucleotide mutation (CCT>CAT) leads proline (A2) to be replaced by histidine (A1) at position 67 in the amino acid sequence of CSN2, which distinguishes the A1 and A2 allelic variations. The substitution of histidine by proline has been reported to avert the enzymatic hydrolysis of the peptide bond between 66th and 67th position and thereby preventing the release of 7 amino acid-bearing (Tyr-Pro-Phe-Pro-Gly-Pro-Ile) bioactive peptide ‘opioid’ is called beta-casomorphin 7 (BCM-7) [10]. In the human small intestine, enzyme digestion leads to the cleavage between histidine and adjacent amino acids successively and releases a BCM-7. Numerous opioid receptors in the human cardiovascular, respiratory, neurological, and immunological systems are potentially affected by BCM-7 [10,11]. A number of human diseases and disorders, including ischemic heart disease, insulin-dependent diabetes, atherosclerosis, schizophrenia, autism, and milk intolerance are thought to be influenced by BCM-7 (A1 variants) [12,13,14]. Several studies, however, have found that cow's milk with a dominating or singular A2-milk is better for human health than A1 milk and poses no threats to health [15,16]. There has been a growing global preference for A2 milk with regard to issues of human health for the last several years. Nowadays, genotyping for CSN2 A1 and A2 variants is routinely carried out in the sire selection process of every renowned breeding service provider globally. Earlier studies reported that most of the Bos taurus dairy breeds possessed A1 CSN2 variants except Jersey and Guernsey which predominantly have A2 variants [17]. On the other hand, the A2 variant is prevalent in different Bos indicus cattle breeds including indigenous cattle of Bangladesh [14,18,19].
The proportion of upgraded Holstein Friesian-Local (HF×L) animals is increasing day by day and there is a possibility of increasing the resultant undesirable A1 allele frequency in the upper graded animals (F2 to onwards). For the genotyping of A1 and A2 allelic variants, a new allele-specific polymerase chain reaction (AS-PCR) method has been developed with newly designed primer sets [19]. Our previous study detected polymorphisms in the coding sequence of CSN2 gene and its allelic variants in indigenous and HF×L crossbred cattle of Bangladesh [19]. To produce milk of the highest quality, it would be beneficial to comprehend the A1 and A2 allelic status of the CSN2 gene in the existing HF×L crossbred cattle population of Bangladesh. However, A1/A2 allele specific genotyping for the aforesaid crossbred cattle population has not yet been performed including diverse sources of samples from private, military and govt. dairy farms. Thus, the aim of this research was to characterize A1/A2 allelic patterns and genotypic frequency using AS-PCR based genotyping in HF×L crossbred cattle population of Bangladesh.
MATERIALS AND METHODS
Ethical statement
Prior approval was taken from the Bangladesh Agricultural University ethical committee (No. 1219/BAURES/2020ESRC/AH/09) for conducting this research and institutional guidelines were followed during experimentation.
Blood sampling
Blood sampling was performed from the HF×L crossbred dairy cattle populations of Bangladesh those were being maintained under intensive management condition at the government dairy farms, military dairy farms and some medium to large sized private dairy farms. Genotypes were ascertained from the recorded pedigree information and in some cases through interaction with farmers. A total of 390 blood samples (3 to 5 ml) were collected from jugular vein using venoject tubes coated with EDTA as an anticoagulant. Blood samples were collected randomly from the unrelated heifers, bulls, and cows. Collected samples were transferred immediately to the Animal Genetics and Genomics lab, Bangladesh Agricultural University, Mymensingh, Bangladesh and preserved at 4-6°C until DNA extraction.
DNA extraction and PCR amplification
DNA was extracted from the whole blood using Add PrepTM Genomic DNA Extraction Kit (ADD BIO Inc., Daejeon, South Korea) according to the manufacturer’s instructions with some modifications. The concentration and purity of isolated DNA were measured by NanoDrop spectrophotometer (Model ND2000, Thermo Fisher Scientific, Wilmington, USA). The A1 and A2 allelic patterns in the HF×L crossbred cattle populations of Bangladesh were investigated in this study using an allele-specific primer that had previously been developed by Pabitra et al. 2022 [19]. Mismatched nucleotides were added to the 3' terminal end of the primers in order to construct them as allele-specific. The primer sequence information is given in Table 1 as previously used [19]. The PCR amplification was performed in a 20 µl reaction volume containing 2 µl DNA, 10 µl of 2× master mix (Prime Taq DNA polymerase 1unit/10 µl, 20 mM Tris-HCl (pH-8.8), 100 mM KCl, 0.2% Triton® X-100, 4.0 mM MgCl2, enzyme stabilizer, sediment, loading dye and 0.5 mM each of dNTP) and 8 µl allele-specific primer mixture (ADD BIO Inc., Daejeon, South Korea) using TGradient Thermocycler (Biometra GmbH, Goettingen, Germany). The thermal profile included an initial denaturation at 95 °C for 10 min, then 30 cycles with denaturation temperature at 95 °C for 20 sec, annealing temperature at 60 °C for 20 sec, extension temperature at 72 °C for 20 sec, and final extension temperature at 72 °C for 5 min.
Table 1. Information on allele-specific primer sequences
Agarose gel electrophoresis and gel documentation
Following PCR, agarose gel electrophoresis was used to verify the amplification of the CSN2 gene sequence fragment. A volume of 20 µl of each PCR product was used for gel electrophoresis in 2.5% agarose gel stained with 5 µl (10 µg/µl) of safe gel dye (green). The results were then observed and documented using the GDS-200 digital gel documentation system (Sunil-Bio, Inc., Seoul, South Korea). The documentation system took images of the gel and annotated them to identify each sample. The images were then analyzed and subsequently genotyped based on the banding pattern. In the A1 and A2 genotyping systems, three distinct band patterns were observed, and thus, the type of genotype was determined.
Statistical analysis
The genotyped data from the gel images was recorded in Microsoft Excel sheets. Genotypic and allelic frequencies at A1/A2 locus were calculated according to Falconer and Mackay [20] by counting the number of bands appearing in the gels. The three genotypes were analyzed by the chi-square test for goodness of fit between observed and expected frequencies with level of significance at probability P<0.05. Descriptive statistics such as mean, percentage and graph were performed using the Microsoft Excel program.
RESULTS
PCR amplification for genotyping of A1 and A2 allelic variants
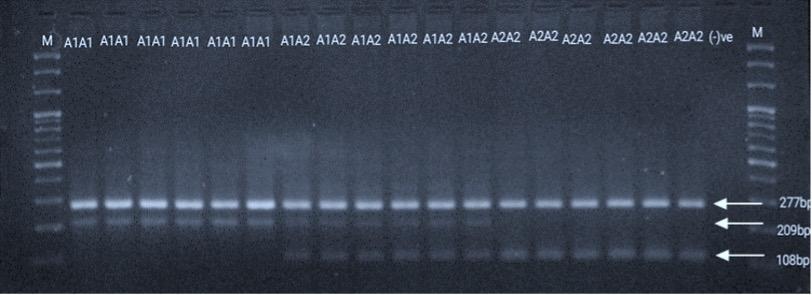
In this study, AS-PCR was used to distinguish A1 and A2 alleles directly from the gel image, based on the 67th amino acid position (Figure 1). In total, 390 crossbred cattle were genotyped based on the three different banding patterns in the gel. All individuals in the AS-PCR genotyping had a common 277 bp fragment, but the allele-specific fragments for the A1A1, A2A2, and A1A2 genotypes were 209 bp, 108 bp, and 209 and 108 bp, respectively (Figure 1). All three genotypes A1A1, A1A2, A2A2 were observed in the investigated samples.
Genotype and allele frequencies of the CSN2 gene variants
Table 2 illustrates the genotype and allele frequencies of the A1 and A2 variants of the CSN2 gene at the farm level in the populations of HF×L crossbred cattle. Both genotype and allele frequencies differed largely among the farms. Altogether, the frequencies of A1A1, A1A2 and A2A2 genotypes were 0.11 (43), 0.48 (187) and 0.41 (162), respectively, whereas the A1 and A2 allele frequencies were 0.34 and 0.66, respectively. In most cases, β-casein gene locus did not follow HW equilibrium at farm level individuals. The undesirable A1A1 genotype frequencies were relatively higher in some private dairy farms that ranged between 6% and 35%. Genotypic and allelic frequencies according to the genetic proportions of HF×L crossbred cattle are shown in Table 3. A total of 81 genotyped information was not included due to absence of recorded pedigree or ambiguity in trace back their genetic proportion properly. In general, A1A1 and A1A2 genotypic frequency increased proportionately as the progression of exotic inheritance level in the crossbred populations. The resultant A1 allele frequency also increased from 24% to 41% in the studied populations. The frequency of desirable A2A2 genotype reduced from 0.60 (50%HF × 50%L) to 0.38 (87.5%HF × 12.5%L) with the advancement of exotic HF inheritance from 50 to 87.5%. The investigated farms were categorized into three groups as government, private and military dairy farms. Farm wise genotypic and allelic frequency is displayed in Table 4. The undesirable A1A1 genotype frequency was found highest in the HF crossbreds of private dairy farms (0.13) compared to government (0.08) and military dairy farm (0.07). The genotype frequencies of heterozygous animals (A1A2) were almost similar in three different farm categories while the A2A2 genotype frequency was significantly lower in private dairy farms.
Table 2. Genotype and allele frequencies of the CSN2 gene based on farm level analysis in Bangladeshi HF×L crossbred cattle herds
Table 3. Genotype and allele frequencies of the CSN2 gene based on genetic proportions of HF×L crossbred cattle of Bangladesh
Table 4. Genotype and allele frequencies of the CSN2 gene in the Bangladeshi HF×L crossbred cattle populations based on farm category
Comparison of the genotypic frequency distribution
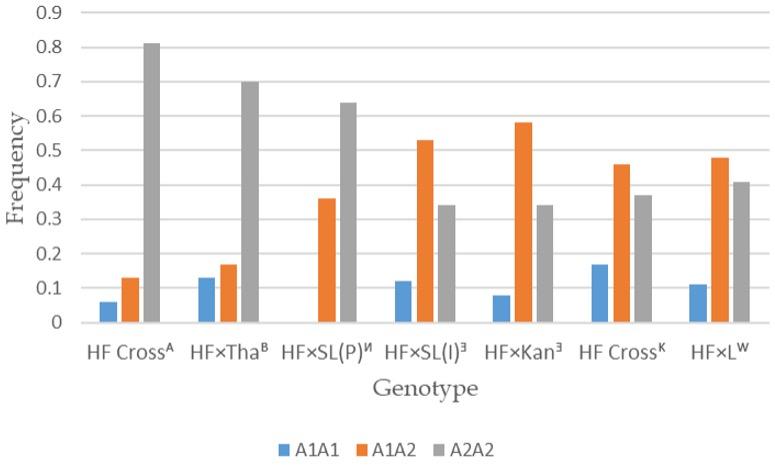
Data on CSN2 genotype frequencies in different Holstein derived crossbred populations were compared and were noticed significant differences among the crossbred cattle populations (Figure 2). The highest A2A2 genotypic frequency was found in Indian HF crossbreds (0.82) and the lowest was in HF×SL (0.34) and HF×Kan (0.34) populations where Bangladeshi HF×L crossbred population occupied the intermediate position (0.41). The highest proportion of heterozygous (A1A2) animals was found in HF×Kan (0.58) population followed by HF×SL (0.52) and HF×L (0.48) crossbred populations. Besides, HF × Kangeyam crossbred displayed the highest genotypic frequency for A1A1 individuals (0.17) and it was absent in one HF×SL population.
DISCUSSION
AS-PCR has been reported at first in cattle by Keating et al. 2008 [21]. Afterwards, this method has been used successfully to genotype A1 and A2 allelic variants in different cattle breeds and populations. Recently, the allele specific PCR amplification was developed by Pabitra et al. 2022 [19] through affiliation of alleles (C to A) in the primer located at the 67th amino acid position in the CSN2 gene's protein coding sequence. In earlier studies, similar AS-PCR approach was used to differentiate between the A1 and A2 allelic variants of CSN2 gene [9, 22, 23] and supports the newly developed protocol by Pabitra et al. 2022 [19]. However, there was basic difference between present and previous studies regarding primer design and the resultant genotyping protocol and allele fragment size.
Chessa et al. 2013 [24] developed two simple tests based on the bi-directional allele-specific polymerase chain reaction (BAS-PCR) to identify genetic variants of β-casein B and I. These tests validate the current AS-PCR approach. Numerous techniques, including PCR-RFLP, PCR-SSCP, AS-PCR, and direct sequencing, have been used to genotype the A1 and A2 allelic variants of the CSN2 gene that are most often found in various cattle populations around the globe [6, 18, 25, 26, 27]. Among these, AS-PCR is one of the most economical and quick methods for determining the genotypes of A1 and A2. The present study demonstrated the efficacy of the newly developed AS-PCR technology in reliably identifying allelic variants. This genotyping protocol has the potential to be economically implemented in many cattle populations worldwide, including the Holstein-derived crossbred cattle population in Bangladesh.
The studied samples of this research had all three genotypes, A1A1, A1A2, and A2A2, with mean genotypic frequencies of 0.11, 0.48, and 0.41, respectively. In line with this result, Kumar et al. 2017 [18] recently found all three types of genotypes in Vrindavani crossbred cattle, with frequencies of 12.3%, 39.6%, and 48.1%, respectively, for A1A1, A2A2, and A1A2 genotypes. According to Ganguly et al. 2013 [22] and Patel et al. 2019 [27], the frequencies of A1A1, A1A2, and A2A2 genotypes were 0.09 vs. 0.15, 0.56 vs. 0.41, and 0.34 vs. 0.44, respectively, in HF×Kankrej and Frieswal (Friesian × Sahiwal) crossbred cattle populations. Moreover, the corresponding A1 and A2 allelic frequencies were 0.38 vs. 0.35 and 0.62 vs. 0.65, respectively, and are nearly identical to the current findings. Our results are corroborated by Miluchová et al. 2014 [28], who found that the proportion of A1A1, A1A2, and A2A2 genotypes in Holstein cattle were 0.14, 0.46, and 0.40, respectively and support our findings. Nonetheless, the A1A1 genotypic proportion in Tamil Nadu's HF was somewhat more than the current findings [9]. In addition, Dai et al. 2016 [6] found that the three genotypes were 0.20 (A1A1), 0.35 (A1A2), and 0.23 (A2A2), respectively, based on direct sequencing of CSN2 gene fragments, which was significantly different from the current study. Altogether, the above stated findings are in agreement with the current result. The significant deviation of HW equilibrium in the studied populations for genotypic frequency might be due to small sample size, continuous selection in the herd and non-random mating. It's worth noting that there isn't much information on taurine × zebu crossbred cattle to compare the findings of this study. Several studies have attempted to study the A1 or A2 milk status where crossbreeding is practiced. Taken together, breed or population-specific gene and genotype frequencies are greatly impacted by a number of variables, including the number of samples investigated, the mating system and genetic drift [20]. Genetic composition of the crossbred animals also had potential influence on CSN2 genotype and allele frequencies [19].
The investigated crossbred animals were grouped into four separate genotypes based on their genetic proportions. In general, the undesirable genotype and allele frequencies were increased proportionately with the increment of exotic inheritance and is consistent with the findings of Jawane et al. 2018 [29] who reported that genotype (A1A1) and gene frequency (A1) was higher in the HF crossbreds having 75% exotic inheritance as compared to 62.5%. They reported overall gene frequencies of A1 and A2 allele in HF crossbreds were 0.08 and 0.92 respectively, whereas it was 0.03 (A1) and 0.97 (A2) in HF 62.5% and 0.13 (A1) and 0.86 (A2) in HF75% crossbreds. However, very limited information is available about the comparative A1/A2 allelic and genotypic frequencies with respect to different exotic inheritance levels in crossbred cattle of Bangladesh and across the world. Importantly, private farms had greater frequency for A1A1, A1A2 and A2A2 genotypes as 0.13, 0.49 and 0.38, respectively compared to government farms where the frequencies of the corresponding genotypes were 0.08, 0.49 and 0.43, respectively. The plausible reasons for higher proportion of A1A1 genotype might be their crossbreds with higher exotic inheritance (75% and 87.5%) for producing higher amount of milk and use of high merit bulls having A1A1 genotype.
CONCLUSION
The total frequencies of A1A1, A1A2 and A2A2 genotypes were 0.11, 0.48 and 0.41 respectively in Bangladeshi HF×L crossbred cattle. The frequencies of A1A1 and A1A2 genotypes were gradually increased with the advancement of exotic Holstein inheritance level. A1A1 genotypic frequency was found highest in 87.5%HF×12.5%L (0.18) crossbreds and was lowest in 50%HF×50%L crossbreds (0.08). In contrast, the A2A2 genotypic frequency declined from 0.60 (50%HF × 50%L) to 0.38 (87.5%HF × 12.5%L) in the studied populations.
In order to decrease the percentage of undesired A1 alleles in crossbred herds, the recently developed AS-PCR technology may be used as an effective marker for screening crossbred cows and breeding bulls for their A1 and A2 allelic variants. This will ultimately result in Bangladeshi crossbred dairy cattle producing milk of higher quality.
ACKNOWLEDGEMENT
This research was carried out with the financial support of a Krishi Gobeshona Foundation (KGF) project, BARC Complex, Bangladesh (Project No. TF 78-L/20). The authors are thankful to government, military, and private dairy farms for providing phenotypic performance data and blood samples.
AUTHOR CONTRIBUTIONS
MAS and MSAB designed of the experiment. MAM and MAS performed the wet lab experimentation and analyzed data. MFAH and SAS were involved in phenotypic data recording and blood sampling. MAS contributed to drafting the article. AKFHB and MSAB contributed to revising it critically and made the final approval of the version to be published.
CONFLICTS OF INTEREST
There is no conflict of interest among the authors.
References
- [1]Farrell HM, Jimenez-Flores R, et al. Nomenclature of the proteins of cows’ milk-sixth revision. J Dairy Sci. 2004; 87: 1641-74.
- [2]Hanusová E, Huba J, et al. Genetic variants of beta-casein in holstein dairy cattle in Slovakia. Slovak J Anim Sci. 2010; 43: 63-66.
- [3]Truswell AS. The A2 milk case: a critical review. Eur J Clin Nutr. 2005; 59: 623-631.
- [4]Boettcher PJ, Caroli A, et al. Effects of casein haplotypes on milk production traits in Italian Holstein and Brown Swiss cattle. J Dairy Sci. 2004; 87: 4311-4317.
- [5]Olenski K, Kamiński S, et al. Polymorphism of the beta-casein gene and its associations with breeding value for production traits of Holstein–Friesian bulls. Livest Sci. 2010; 131: 137-140.
- [6]Dai R, Fang Y, et al. Identification of alleles and genotypes of beta-casein with DNA sequencing analysis in Chinese Holstein cow. J Dairy Res. 2016; 83: 312-6.
- [7]Meier S, Korkuc P, et al. DNA sequence variants and protein haplotypes of casein gene in German Black Pied Cattle (DNS). Front Genet. 2019; 10: 1129.
- [8]Singh LV, Jayakumar S, et al. Comparative screening of single nucleotide polymorphisms in β- casein and κ-casein gene in different livestock breeds of India. Meta Gene. 2015; 4: 85-91.
- [9]Malarmathi M, Kumar TMAS, et al. Analysis of β-casein gene for A1 and A2 genotype using allele specific PCR in Kangeyam and Holstein Friesian crossbred cattle in Tamil Nadu. Indian J Vet Anim Sci Res. 2014; 10: 310-315.
- [10]Bell SJ, Grochoski GT, et al. Health implications of milk containing beta-casein with the A2 genetic variant. Crit Rev Food Sci Nutr. 2006; 46: 93-100.
- [11]Gill SH, Doull F, et al. Immunoregulatory peptides in bovine milk. Br J Nutr. 2000; 84: 111-117.
- [12]Kamiński S, Ruść A, et al. A note on frequency of A1 and A2 variants of bovine beta-casein locus in Polish Holstein bulls. J Anim Feed Sci. 2006; 15:195-98.
- [13]Caroli AM, Chessa S, et al. Invited review: milk protein polymorphisms in cattle: effect on animal breeding and human nutrition. J Dairy Res. 2009; 92: 5335-52.
- [14]Mishra BP, Mukesh M, et al. Status of milk protein, β-casein variants among Indian milch animals. Indi J Anim Sci. 2009; 79: 722-25.
- [15]McLachlan CN. β-casein A1, ischaemic heart disease mortality and other illnesses. Med Hypotheses 2001; 56: 262–272.
- [16]Kaskous S. A1 and A2 milk and their effect on human health. Journal Food Eng Technol. 2020; 9: 15-21.
- [17]Cieślińska A, Kamiński S, et al. Beta-casomorphin 7 in raw and hydrolyzed milk derived from cows of alternative β-casein genotypes. Milchwissenschaft 2007; 62: 125-127.
- [18]Kumar S, Singh RV, et al. Molecular characterization of A1/A2 Beta-casein Alleles in Vrindavani crossbred and Sahiwal cattle. Indian J Anim Res. 2017; 53: 151-55.
- [19]Pabitra MH, Maruf TM, et al. Molecular characterization and A1/A2 genotyping of casein beta gene in zebu and crossbred cattle of Bangladesh. Czech J Anim Sci. 2022; 67: 318-326.
- [20]Falconer DS, Mackay TFC. Introduction to Quantitative Genetics. Essex, England: Longman.: 1996.
- [21]Keating AF, Smith TJ, et al. A note on the evaluation of a beta-casein variant in bovine breeds by allele-specific PCR and relevance to casomorphin. Irish Journal Agril Food Res. 2008; 47: 99-104.
- [22]Ganguly I, Kumar S, et al. Status of β-casein (CSN2) polymorphism in Frieswal (HF X Sahiwal Crossbred) cattle. Int J Biotec Bioeng Res. 2013; 4: 249-56.
- [23]Ullah S, Ahmed Z, et al. Genetic Association of CSN2 variant 6:85451298C>A with A1/A2 milk phenotype in Pakistani cows. bioRxiv 2022; 2022-08.
- [24]Chessa S, Bulgari O, et al. Bovine β-casein: Detection of two single nucleotide polymorphisms by bidirectional allele specific polymerase chain reaction (BAS-PCR) and monitoring of their variation. Open J Anim Sci. 2013; 3: 36-41.
- [25]Massella E, Piva S, et al. Evaluation of bovine beta casein polymorphism in two dairy farms located in northern Italy. Italian J Food Safety. 2017; 6: 6904
- [26]Mclachlan CNS. Breeding and milking cows for milk free of β-casein A1. U.S. Patent No. 7,094,949. Washington, DC: U.S. Patent and Trademark Office, 2006.
- [27]Patel AC, Patel C, et al. Genotyping of Indigenous cattle breeds and their exotic crosses for β -Casein milk type using PCR-RFLP and sequencing. Int J Livest Res. 2019; 9: 249-59.
- [28]Miluchová M, Gábor M, et al. Analysis of Beta-Casein gene (CSN2) polymorphism in different breeds of cattle. Sci Papers Anim Sci Biotechnol. 2014; 47: 56–59.
- [29]Jawane VB, Ali SS, et al. Genetic polymorphism of β-casein (CSN2) in Zebu and HF crossbreds. Indian J Dairy Sci. 2018; 71: 530-533.