Impact of single nucleotide polymorphisms of immune checkpoint CTLA-4 (SNP rs231775 and rs5742909) in susceptibility to Hashimoto's thyroiditis patients
Abstract
Hashimoto thyroiditis (HT) is an autoimmune disorder causing thyroid cell destruction. This research aimed to evaluate CTLA-4 expression levels in patients with HT. As well as understand the risk and protective effects of two CTLA-4 SNPs polymorphisms (rs231775 and rs5742909) in Iraqi patients. A case control study was conducted on 50 HT patients (5 men and 45 females) which were assembled from Baghdad teaching hospital, Baghdad, Iraq. The relative gene expression method was used to measure the gene expression of CTLA-4 using qRT-PCR. The single nucleotide polymorphisms of (318C/T (rs231775) and +49A/G (rs5742909) were sequenced for data analysis. There was a significant increase of serum CTLA-4 levels and, there was a significant increase in fold change of CTLA-4 in HT patients compared to controls. The allele and genotype frequencies according to Hardy Weinberg equilibrium showed significant differences in HT patients in the study, the 318C/T (rs231775) variant showed differences in the AA genotype when compared to controls. However, there were no significant differences observed in the AG and GG genotypes. Similarly, for the CTLA-4 SNP +49A/G (rs5742909), no significant differences were found in the CC, CT, and TT genotypes. Despite these observations, both studied SNPs showed an increased expression of the soluble form and fold change of the immune checkpoint CTLA-4 in HT patients. The study reveals that CTLA-4 gene SNPs rs231775 and rs5742909 significantly influence the expression of CTLA-4 levels in HT patients, which potentially may cause HT incidence and development.
INTRODUCTION
Two to five percent of the global population suffers with Hashimoto's thyroiditis (HT), also known as chronic lymphocytic thyroiditis or autoimmune thyroiditis (AIT). Depression, metabolic syndrome, osteoporosis, and infertility or premature birth in women are all linked to the aberrant thyroid function [1]. It is estimated that genetic factors are responsible for 70% of susceptibilities to disease, and there is a complex interaction between genetic and environmental factors that create autoimmune thyroid disorders (AITD). Whereas the environmental factors include smoking, alcohol, stress, infection, and intake dose of vitamin D, selenium, and iodine [2].
One of the most important factors in regulating T cell activation is a transmembrane protein encoded by the cytotoxic T lymphocyte-associated antigen 4 (CTLA-4). The gene for the transmembrane glycoprotein CTLA-4, also known as CD152, encodes 223 amino acids. Hemostasis and negative control of immune responses rely on CTLA-4 protein, which is expressed on the surface of activated T cells [3]. CTLA-4 gene is expressed on activated T CD4+ and T CD8+ cells [4]. To this end, it negatively regulates T cell activity and contributes to several immune responses such as ensuring peripheral tolerance persists, preventing the production of inflammatory cytokines, and prompting death in activated T cells. It is possible that CTLA-4 regulates the signals that cause T cells to differentiate into Th1 or Th2 by acting on their activation pathways [5]. Polymorphisms in the CTLA-4 gene have been linked to the development of various autoimmune disorders [6]. Variations in the CTLA-4 gene, known as polymorphisms, are strongly linked to autoimmune disorders. These polymorphisms include features like promoters, microsatellite (AT)n repeats found in the 3'-untranslated regions, and changes related to methylation and acetylation [7, 8].
The polymorphisms 318C/T (rs5742909) and +49A/G (rs231775) in the CTLA-4 gene are among the most studied [9, 10]. Immune tolerance requires the presence of the transcription factors such as Aire and Fezf2, which control the production of tissue-restricted antigens in medullary thymic epithelial cells (mTEC). Each transcription regulator uses a unique set of methods to regulate the gene expression of CTLA-4 [11]. During T lymphocyte activation, antigen presented by antigen presenting cells (APC) is recognized by T lymphocytes expressing the T cell receptor (TCR). For T cell activation, further stimulation from the interaction of CD28 and CD80/86 receptor is required. Without a co-stimulatory signal, T cells become unresponsive and anergic [12].
Signals suppressing T cell activity are generated by the interaction of CTLA-4 and PD-1 receptors (on T cell surface) with their ligands (CD80/86 and PD-L1/2). By activating several mechanisms, T-regs suppress the activity of T CD4+ and T CD8+ cells, including the secretion of cytokines interleukin-10 (IL-10), transforming growth factor b (TGF-b) and IL-35 inhibits T cell growth, IL-2 causes T cell death, and granzymes harm T cells. Direct action by Tregs is possible via surface-expressed molecules that bind to T cell surface receptors and inhibit proliferation [13]. CTLA-4 suppresses immunological responses by interfering with CD28 co-stimulatory molecules' ability to attach to B7-1 and B7-2 molecules on APC [14]. CTLA-4 and CD28 connect to B7 molecules, influencing the type of immune response that occurs. As a result, variables influencing the expression or activation of CTLA-4 can disrupt this balance, leading to a loss of control over immune responses and, ultimately, triggers autoimmunity [15-17].
Variations in the CTLA-4 gene affect its functionality, contributing to the onset of autoimmune diseases [18]. Studies have examined the association of +49A/G in with rs5742909 and rs231775 in HT [19]. During negative selection, mTEC presents peptides-major histocompatibility complex (MHC) to T cells which helps in eliminating autoreactive lymphocytes [20]. Lymphocyte anergy refers to its inability to respond to an antigen [21]. CTLA-4 and PD-1 receptors, when interacting with APCs expressing CD80/86 and PD-L1/2, inhibit T cells [22].
The increase of APCs expressing MHC class II molecules has been related to genetic variation in CTLA-4, PD1, and other immunological checkpoints. The infiltration of APC (especially dendritic cells and macrophages) into the thyroid may be triggered by inflammation caused by a viral or bacterial infection or by the exposure of thyroid cells to toxins. Thyroid follicular cells from AITD patients also produce elevated levels of interferon-gamma (IFNG)-induced MHC class II molecules, which aid in the presentation of thyroid autoantigens and the activation of T lymphocytes. Up to half of the infiltrating immune cells in advanced thyroiditis are B cells [23]. Other invading immune cell types include cytotoxic T lymphocytes and CD4+ cells [24].
T CD4+ cells are activated upon contact with APC, and further division results in the development of Tregs and Th lymphocytes (including Th1, Th2, and Th17 cells) [25]. Thyroxine (Th) and thyroxine (Tc) induce many processes including cytokines and/or cytotoxins that lead to the death of the thyroid gland in HT. Tregs, which usually suppress pro-inflammatory Th17 activity, are diminished in people with AITD. Loss of self-tolerance in autoimmune disorders is thought to be caused by two main mechanisms: pro-inflammatory Th17 activation and Tregs attenuation [26]. This study aimed to explore the link between the 318C/T (rs5742909) and +49A/G (rs231775) SNPs, CTLA-4 gene expression, and immunological intolerance due to the limited findings in previous research.
MATERIALS AND METHODS
Subjects
The study registered HT patients, who were advised to hospital and used therapy, including biotherapy, NSAID, or SAID as inclusion criteria. 50 patients aged 13-65 were gathered from Baghdad teaching hospital, Iraq. Patients were sub-grouped into male and female, where the exclusion criteria were those with diabetes type 1 and type 2 and additional autoimmune illnesses. For the purpose of comparison 50 healthy subjects were also registered in the current study as case control.
The Institutional Review Board approved experimental research involving humans or animals (Ministry of Higher Education and Scientific Research, Al-Nahrain University, Scientific Research Ethics Committee, number 164, Dated 03-09-2022) before conducting research and collecting samples. The investigation did not contain any special pictures of patients, healthy people, or any part of them to protect the patient's identity and privacy.
Sample collection
From each participating patient, 10 ml of venous blood was collected under aseptic conditions from the patients and the controls. The blood was distributed into aliquots (3 ml) in EDTA tubes. The aliquots were processed after reaching the laboratory to isolate total DNA for SNP analysis. And 300 ul of whole blood was add to 600 ul Trizol reagent (TransGene, China). While the remaining 5 ml aliquot was dispensed in gel tube for serum separation for laboratory investigations.
Extraction of total DNA and RNA
ReliaPrep™ DNA and RNA extraction Kit (Promega, USA) was used for extraction and purification of DNA from blood samples, and the methods were carried out according to the instruction manual supplied. The concentrations of DNA and RNA were estimated using QuantiFluor®, Promega, USA.
Primers were used in this study.
The primer sequences for human Rs231775 CTLA-4 SNP were forward primer 5' ggcttgccttggatttcagc-3', and reverse primer 5'aaaacaggagagtgcagggc-3' (296 bp). The primer sequences for human rs5742909 SNP were forward primer 5' ccttgtactccaggaaattctccaa-3', and reverse primer 5' gaagcttcatgttcactttgagg-3' (231 bp). The primer sequences for human CTLA-4 were forward primer 5’-ctacctgggcataggcaacg-3’, and reverse: 5’- ccccgaactaactgctgcaa -3’. These primers were created and used according to the producer Takara Bio Inc. (Shiga, Japan). The human glyceraldehyde-3-phosphate dehydrogenase (GAPDH) was used as endogenous control, and the forward primer was 5'gacagtcagccgcatcttct3', and the reverse primer was 5'gcgcccaatacgaccaaatc3' (Takara Bio Inc., Shiga, Japan).
Determining the expression of CTLA-4
Following the manufacturer's instructions, human CTLA4 ELISA Kit was used to estimate the concentration of CTLA-4 in HT and control (MyBioSource, USA). Following the manufacturer's instructions, RNA was extracted and purified from blood samples using Relia-PrepTM RNA Miniprep (Promega, USA). England's Reverse Transcriptase (RT) Mix Reagent the SuperScript IV-VILO Master Mix from Invitrogen, USA was used to transcribe total RNA into complementary DNA. A SaCycler-48 thermal cycler from Sacace, Italy, was used to carry out the cDNA preparation. KAPA-SYBR® Fast qPCR master mix, USA was used to make the reaction mixture for quantitative PCR (qPCR). As an endogenous control, GAPDH was used as housekeeping gene. The topographic separation of double-stranded DNA (dsDNA) throughout cycles of increasing denaturing temperature allowed for its melting-curve characterization.
Statistical analysis
Graph Pad Prism v8 and IBM SPSS Statistics v27 were used to calculate the mean and standard error, and correlation, where p=0.05 was deemed non-significant. Using the equations CT = CT of target gene - CT of U gene, CT = CT of each sample - average control C, and Fold change = 2-Ct, the fold change was calculated. It was noticed that the control value was regarded to be 1, samples with values less than 1 are considered down-regulated, while samples with values greater than 1 are considered up-regulated 10. Online calculator https://wpcalc.com/en/equilibrium-hardy-weinberg was used to determine allele frequencies. The H-W calculator for two alleles, which can be found for free at www.dr-petrek.eu/documents/HWE.xls, was used to estimate a considerable divergence from Hardy-Weinberg equilibrium (HWE). The distribution of SNP genotypes was reported as a percentage, and statistical significance between gout patients and controls was determined using Fisher's exact probability test (p). Odds ratios (OR) and 95% confidence intervals were calculated to determine the risk connection between the genotype and gout susceptibility (95 percent CI). An OR number below one indicates a negative relationship, whereas an OR value over one indicates a positive one. These estimates were determined with the use of the epidemiologist oriented WINPEPI software. The most up-to-date version of WINPEPI (containing all applications and documentation) may be downloaded for free from http://www.brixtonhealth.com.
RESULTS
Distribution of HT patients
The age of patients ranged from 22 to 68. Table 1 showed that females were much more likely to be affected by HT (90% vs. 10%). The rates of HT patients were significantly higher (84% vs. 16%) among those aged 30 and older than other age groups (Table 1).
Table 1. Distribution of HT patients according to gender and age.
Role of single nucleotide polymorphism CTLA-4 SNP rs 231775 in HT
The human SNP of CTLA4 gene (rs231775 A/G), that located in 2q33 at position +49 of exon-1 on chromosome 2 represented three genotypes (AA, AG, and GG) that are corresponding to two alleles (A and G). The study found that HT patients had three genotypes AA, AG, and GG, which accounted for 42%, 50%, and 8% of their genotypes, respectively, compared to 18%, 70%, and 12% of healthy controls (Table 2). These genotypes showed a strong agreement with HWE, suggesting protected variants. The AA genotype showed significant differences, while no significant differences were found in other genotypes. The allele frequencies of the A and G alleles also showed significant differences (Table 2).
Table 2. Distribution of the observed rs231775 genotype and allele frequencies in HT patients.
Role of single nucleotide polymorphism CTLA-4 SNP rs5742909 in HT
The study found that the three genotypes such as CC, CT, and TT in HT patients and controls were similar, with no significant differences in observed and expected genotype frequencies. The allele frequencies of the C and T alleles showed no significant differences between HT patients and controls. The genotypes showed a strong agreement with HWE (Table 3).
Table 3. Distribution of the observed rs5742909 genotype and allele frequencies in HT patients.
Effect of CTLA-4 rs 231775 SNP on expression of CTLA-4 levels in HT patients
The study found significant increases in serum CTLA-4 levels in HT patients, where genotypes AA and AG and GG showing significant increases in CTLA-4 levels compared to the control group (Figure 1). The CTLA-4 gene fold change was significantly higher in AA, AG, and GG genotypes compared to the control group (Figure 2).
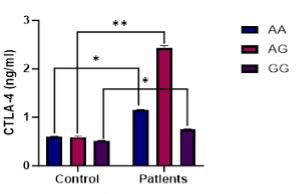

Effect of CTLA-4 rsrs5742909 SNP on expression of CTLA-4 in HT patients
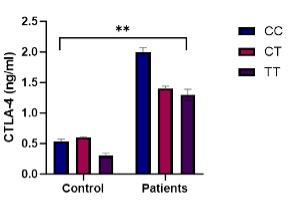
The study found a significant increase in serum CTLA-4 levels in HT patients compared to the control group, where CC genotypes showing a higher sCTLA-4 than CT and TT genotypes (Figure 3). The CTLA-4 gene fold change was significantly higher in CT, and CC genotypes and no significant differences recorded in TT genotype in HT patients compared to the controls (Figure 4).

DISCUSSION
Considering the ability of CTLA-4 to dampen the activity of the immune system, the variables that affect its expression and function may be considered pivotal in immunological regulation. SNPs may not be directly responsible for disease, but they are still a major risk factor because of their impact on gene expression and the products of that gene. A common amino acid change, from Thr to Ala, is the result of the +49A/G (rs231775) polymorphism in exon 1 of the CTLA-4 gene. This defect prevents the CTLA-4 protein from performing its regular duties in the endoplasmic reticulum. This SNP reduces CTLA-4 protein glycosylation, which in turn reduces CTLA-4 protein synthesis at the cell surface [1]. Reductions in T-cell proliferation have been associated with the G allele. A meta-analysis found that the CTLA-4 gene was linked to the risk of HT, and that those who carried both the dominant G allele (GG + GA) and the recessive allele (GG) of this polymorphism were at a higher risk of developing HT, while a meta-analysis study did not find this association [27]. Numerous prior investigations support our findings and reveal that mutations in the CTLA-4 gene, which is located on chromosome 2q33, might lead to a reduced inhibitory impact on the T cell response. Due to its association with a wide range of systemic and autoimmune disorders, including insulin-dependent diabetic mellitus, rheumatoid arthritis (RA), and HT, the present study investigated the functional SNP rs231775 in exon 1 of the CTLA-4 gene. In addition, the rs231775 GG genotype is more common in Caucasian individuals with RA and HT and has been linked to enhanced T cell activation and proliferation [28].
The meta-analysis included 2,315 RA patients and 2,536 controls who were selected using CTLA4 exon1 A49G genotyping from 10 studies [29]. A study also reported the increase of CTLA-4 in HT and the incidence of autoimmune thyroid disease through the dysregulation of immune tolerance [30]. Different hypotheses on the role of the CTLA4 exon1 A49G polymorphism in RA susceptibility were found in subgroup and meta-regression analyses depending on ethnicity (European vs. Asian) [31, 32]. No association between G and susceptibility was found in people of European ancestry. The CTLA4 exon1 49G allele is not associated with increased risk of RA or HT in Europeans, but it may have a role in RA susceptibility in Asians, as shown by a meta-analysis. The distribution of A49G genotypes and alleles is not significantly different between RA patients and controls, which is consistent with our findings (Tables 1 and 2). There was also a significant increase in the frequency of the G/G genotype of CTLA4 A49G polymorphism in HT patients compared to controls, although this difference did not reach statistical significance in many papers because of the limited sample size of HT patients [33]. In general, genotype and allele frequencies of A49G polymorphism were supposed to have direct effects on the elevated level of CTLA-4 and could be risk factor for the development of HT for the allele genotype A while the G allele may have a protected effects on the developing HT [34]. CTLA4 gene +49A/G polymorphisms associated with the AA genotype may increase the risk of BD by interfering with immunosuppression via the CTLA-4 signaling pathway. The mean blood level of sCTLA-4 was lowered in BD patients, particularly in those with the CTLA4 +49 G allele [35]. However, the A allele in patients was the genetic component responsible for the decrease in CTLA-4 level [35]. T cells with abnormal CTLA-4 expression are susceptible to immunological homoeostasis imbalance. CTLA-4 +49 G/A rs231775 is one of the most well investigated genetic variables in genome research [36].
Previous studies have shown that a rs5742909 polymorphism, whereby the C allele is changed to the T, increases CLTA-4 protein synthesis, and may lower the risk of numerous autoimmune illnesses. This study indicates that HT patients had a greater frequency of T alleles than healthy people, where the inverse was true for C alleles [37]. This data raises the possibility that the CTLA4 C allele protects against HT and that the CTLA4 T allele is an up-regulated risk factor for its development [37]. It is known that the transition from the C (wild type) allele to the T (mutant) allele in the rs5742909 polymorphism increases the expression of CTLA-4 proteins [38]. The increased CTLA-4 expression in peripheral blood may reflect the continuous T cell activation in chronic infections and autoimmune disorders. Similarly, CTLA-4 expression is high in RA, lupus erythematosus, and HT. However, the reason of T cell activation is unknown. Thus, the increased expression of CTLA-4 may explain a role in the formation of Th1 cells, the most common subset of T cells, which are considered to drive harmful effector pathways in autoimmune disorders. In contrast, it is known that CTLA-4 plays a role in the formation and maintenance of peripheral immunological tolerance following CTLA-4 inhibition. Therefore, CTLA-4 may similarly prevent chronic T cell activation [39]. The relationship between the 318C>T polymorphism and other autoimmune disorders, including spondyloarthropathy, pemphigus foliaceus, multiple sclerosis, Behcet's disease, systemic lupus erythematosus, HT, ankylosing spondylitis, and Graves' disease, supports our findings [40]. Despite the higher C allele in controls and the increased T allele in HT patients, the -318C/T polymorphism was not significantly associated with HT in our case-control study. A study on polymorphism and CTLA4 protein expression showed that the -318T allele has higher promoter activity. No significant differences were observed, aligning with our results. This consistency might be due to the small sample size used [41]. Evidence suggests that changes to the promoter region might affect gene expression. The hypothesis indicates that the 318C/T (rs5742909) SNP in the CTLA-4 gene affects promoter activity, which in turn influences the amount of CTLA-4 expressed on the cell surface [41]. Increased CTLA-4 mRNA expression was seen in T allele carriers, which was followed by increased CTLA-4 protein expression on the cell surface. While some research has associated the CTLA-4 +49A/G SNP with autoimmune diseases like RA in Mexico, other studies have not found a connection between CTLA-4 SNPs and autoimmune thyroid disease or any other autoimmune conditions in different populations [42].
Numerous research has proposed several possible explanations for these divergent findings. The ethnic characteristics of the study populations are varied, the study populations range in size, and the autoimmune diseases investigated are themselves complex. The 318C>T polymorphism, the +49A>G polymorphism, and the CT60A>G polymorphism are reported because they affect promoter activity and transcription efficiency (318C>T), T cell activation through posttranslational modification (+49A>G), and alternative splicing and CTLA-4 isoform production (CT60A>G), respectively, thus can affect autoimmune disease and pathogenesis. Despite the modest size of samples of the current study, it is certain that the current findings will add to the growing body of meta-analysis research attempting to pin down CTLA-4's role in HT's pathogenesis [43].
CONCLUSIONS
The study found that CTLA-4 gene SNP significantly influences soluble CTLA-4 levels in rs231775 HT patients, with increased AA genotype in frequency potentially causing Hashimoto disease and lower CC and higher CT genotypes could be related to the incidence of HT. Also, rs5742909 demonstrates non-significant differences in the alleles and genotypes frequencies.
ACKNOWLEDGEMENT
The authors are grateful to the Central Blood Bank in Baghdad and Baghdad Teaching Hospital for their cooperation during the collection of blood samples from patients.
AUTHOR CONTRIBUTIONS
AMM and YWI planned, designed outlines, collection, preparation, working procedures, data collection and analysis in the manuscripts. RHA performed experimental design, supervision, and article writing. All authors approved final version of the manuscript.
CONFLICTS OF INTEREST
There is no conflict of interest among the authors.
References
- [1]Kaykhaei MA, Moghadam H, et al. Association of CTLA4 (rs4553808) and PTPN22 (rs2476601) gene polymorphisms with Hashimoto’s thyroiditis disease: A case-control study and an In-silico analysis. Meta Gene. 2020; 24:100693.
- [2]Issa YW, Salih SM. Cytokines, and oxidative stress profiles in Iraqi patients with vitiligo. Ann Trop Med Public Health. 2019;22(8).
- [3]Borysewicz-Sańczyk H, Sawicka B, et al. Genetic Association Study of IL2RA, IFIH1, and CTLA-4 Polymorphisms with Autoimmune Thyroid Diseases and Type 1 Diabetes. Front Pediatr. 2020; 8:481.
- [4]Ramgopal S, Rathika C, et al. Interaction of HLA-DRB1* alleles and CTLA4 (+49 AG) gene polymorphism in Autoimmune Thyroid Disease. Gene. 2018; 642:430–8.
- [5]Harrison SR, Li D, et al. Vitamin D, Autoimmune Disease and Rheumatoid Arthritis. Calcif Tissue Int. 2020;106(1):58.
- [6]Joob B, Wiwanitkit V. CTLA4 polymorphisms and Graves’ disease. Cell Immunol. 2020;348.
- [7]T Onoue DUNBYTHYMS. Epithelial-mesenchymal transition induced by the stromal cell-derived factor-1/CXCR4 system in oral squamous cell carcinoma cells. Int J Oncol. 2006; 29:1133–8.
- [8]Ono K, Ide K, Kashihara M, et al. Polymorphisms in CTLA-4 predict de novo donor specific antibody formation after kidney transplantation. Hum Immunol. 2022;83(6):494–8.
- [9]Abbaspour-Aghdam S, Ahmadi M, et al. Molecular analysis of CTLA4 gene in patients with Behçet’s disease from an Iranian Northwest Azeri population. Gene Rep. 2020; 19:100612.
- [10]Ueda H, Howson JMM, et al. Association of the T-cell regulatory gene CTLA4 with susceptibility to autoimmune disease. Nature. 2003;423(6939):506–11.
- [11]Chikuma S. CTLA-4, an essential immune-checkpoint for T-cell activation. In: Emerging Concepts Targeting Immune Checkpoints in Cancer and Autoimmunity. Springer; 2017. p. 99–126.
- [12]Tokić S, Ŝtefanić M, et al. Altered expression of CTLA-4, CD28, VDR, and CD45 mRNA in T cells of patients with Hashimoto’s thyroiditis - a pilot study. Endokrynol Pol. 2017;68(3):274–82.
- [13]Williams DE, Le SN, et al. Thyroid Peroxidase as an Autoantigen in Hashimoto’s Disease: Structure, Function, and Antigenicity. Horm Metab Res. 2018;50(12):908–21.
- [14]Krysiak R, Kowalcze K, et al. Selenomethionine potentiates the impact of vitamin D on thyroid autoimmunity in euthyroid women with Hashimoto’s thyroiditis and low vitamin D status. Pharmacological Reports. 2019;71(2):367–73.
- [15]R Zhao HZJZXLSB. Interleukin-1β promotes the induction of retinal autoimmune disease. Int Immunopharmacol. 2014; 22:285–92.
- [16]H Wang Y. Development of an inactivated vaccine candidate, BBIBP-CorV, with potent protection against SARS-CoV-2. Cell. 2020;182(3):713 e719-721 e719.
- [17]Liu W, Yang Z, et al. The Association Between CTLA-4, CD80/86, and CD28 Gene Polymorphisms and Rheumatoid Arthritis: An Original Study and Meta-Analysis. Front Med (Lausanne). 2021 Feb 2; 8:598076.
- [18]Narooie Nejad M, Taji O, et al. Association of CTLA 4 gene polymorphisms 318C/T and +49A/G and hashimoto’s thyroidits in Zahedan, Iran. Biomed Rep. 2017;6(1):108–12.
- [19]Fathima N, Sultana Q, et al. Autoimmune thyroid patients with CTLA-4 (+49A/G) GG/AG genotypes have high seropositivity to thyroid peroxidase than thyroglobulin. Meta Gene. 2022; 31:101010.
- [20]Benvenga S, Guarneri F. Homology between TSH-R/Tg/TPO and Hashimoto’s encephalopathy autoantigens. Frontiers in Bioscience - Landmark. 2020;25(2):229–41.
- [21]Ahmed R, Al-Shaikh S, et al. Hashimoto Thyroiditis. Adv Anat Pathol. 2022;19(3):181–6.
- [22]Godlewska M, Banga PJ. Thyroid peroxidase as a dual active site enzyme: Focus on biosynthesis, hormonogenesis and thyroid disorders of autoimmunity and cancer. Biochimie. 2019; 160:34–45.
- [23]Klubo-Gwiezdzinska J, Wartofsky L. Hashimoto thyroiditis: an evidence-based guide to etiology, diagnosis, and treatment. Pol Arch Intern Med. 2022;132(3).
- [24]Kaykhaei MA, Moghadam H, et al. Association of CTLA4 (rs4553808) and PTPN22 (rs2476601) gene polymorphisms with Hashimoto’s thyroiditis disease: A case-control study and an In-silico analysis. Meta Gene. 2020; 24:100693.
- [25]Capriello S, Ferrari SM, et al. Regulatory B Cells in Systemic Sclerosis Isolated or Concomitant with Hashimoto Thyroiditis. Front Immunol. 2022;13.
- [26]Taheri M, Eghtedarian R, et al. Dysregulation of non-coding RNAs in autoimmune thyroid disease. Exp Mol Pathol. 2020; 117:104527.
- [27]Lefort S, Thuleau A, et al. CXCR4 inhibitors could benefit to HER2 but not to triple-negative breast cancer patients. Oncogene. 2017;36(9):1211–22.
- [28]Boomer JS, Green JM, et al. The changing immune system in sepsis is individualized immuno-modulatory therapy the answer virulence. 2014;5(1):45–56.
- [29]Klubo-Gwiezdzinska J, Wartofsky L. Hashimoto thyroiditis: an evidence-based guide to etiology, diagnosis and treatment. Pol Arch Intern Med. 2022;132(3).
- [30]Skov J, Kuja-Halkola R, et al. Shared etiology of type 1 diabetes and Hashimoto’s thyroiditis: a population-based twin study. Eur J Endocrinol. 2022;187(1):677–85.
- [31]Uhliarova B, Hajtman A. Hashimoto’s thyroiditis - an independent risk factor for papillary carcinoma. Braz J Otorhinolaryngol. 2018;84(6):729–35.
- [32]Narooie Nejad M, Taji O, et al. Association of CTLA 4 gene polymorphisms 318C/T and +49A/G and hashimoto’s thyroidits in Zahedan, Iran. Biomed Rep. 2017;6(1):108–12.
- [33]Benhatchi K, Jochmanová I, et al. CTLA4 exon1 A49G polymorphism in Slovak patients with rheumatoid arthritis and Hashimoto thyroiditis-results and the review of the literature. Clin Rheumatol. 2011;30(10):1319–24.
- [34]Narooie Nejad M, Taji O, et al. Association of CTLA-4 gene polymorphisms −318C/T and +49A/G and Hashimoto’s thyroidits in Zahedan, Iran. Biomed Rep. 2017;6(1):108.
- [35]Pastuszak-Lewandoska D, Sewerynek E, et al. CTLA-4 gene polymorphisms and their influence on predisposition to autoimmune thyroid diseases (Graves’ disease and Hashimoto’s thyroiditis). Arch Med Sci. 2012;8(3):415.
- [36]Lee HJ, Li CW, et al. Immunogenetics of autoimmune thyroid diseases: A comprehensive review. J Autoimmune. 2015; 64:82–90.
- [37]Xue H, Yu X, et al. The possible role of CD4+CD25highFoxp3+/CD4+IL-17A+ cell imbalance in the autoimmunity of patients with Hashimoto thyroiditis. Endocrine. 2015;50(3):665–73.
- [38]Krupińska J, Urbanowicz W, et al. Association between genetic mutations and the development of autoimmune thyroiditis in patients with chronic hepatitis C treated with interferon alpha. Thyroid Res. 2012;5(1):1–8.
- [39]Gunavathy N, Asirvatham A, et al. Association of CTLA-4 and CD28 Gene Polymorphisms with Type 1 Diabetes in South Indian Population. 2019;48(6):659–71.
- [40]Hu Y, Xu K, et al. Associations Between Three CTLA-4 Polymorphisms and Hashimoto’s Thyroiditis Risk: An Updated Meta-Analysis with Trial Sequential Analysis. Genet Test Mol Biomarkers. 2018;22(4):224–36.
- [41]Indhumathi S, Rajappa M, et al. T helper-2 cytokine/regulatory T-cell gene polymorphisms and their relationship with risk of psoriasis in a South Indian Tamil cohort. Hum Immunol. 2017;78(2):209–15.
- [42]Soskic B, Qureshi OS, et al. A transendocytosis perspective on the CD28/CTLA-4 pathway. Adv Immunol. 2014; 124:95–136.
- [43]Rowshanravan B, Halliday N, et al. CTLA-4: a moving target in immunotherapy. Blood. 2018;131(1):58–67.